Introducing the Nalo Meli ‘Āpa‘akuma project: Using Genomic Data to Inform Yellow-faced Bee Managemement and Conservation
Jonathan B. Koch, Qindasi Silva, Cynthia King, and Jolene Sutton | http://jonathanbkoch.weebly.com

Abstract
- nalo meli ʻāpaʻakuma translates from ʻōlelo Hawaiʻi as endemic bee.
- Hawaiʻi houses >60 nalo meli ʻāpaʻakuma - bees found nowhere else.
- 7 Hylaeus bees are protected under the U.S. Endangered Species Act.
- The goals of the nalo meli ‘āpa‘akuma project are to generate a genomic resource and population genomic data.
- The data generated will be useful to the stakeholders when they draft and execute a recovery plan for the endangered Hawaiian Hylaeus.

Project Objectives
- Objective 1 | Investigate non-lethal DNA sampling strategies. Examine utility of non-lethal genomic sampling methods of endangered and non-endangered Hylaeus bees.
- Objective 2 | Sequence and assemble genome of endangered H. anthracinus (Figure 1).
- Objective 3 | Characterize population genomic diversity. Test for the effect of habitat fragmentation on population genomic diversity of non-Endangered and Endangered Hylaeus species.

Project Updates
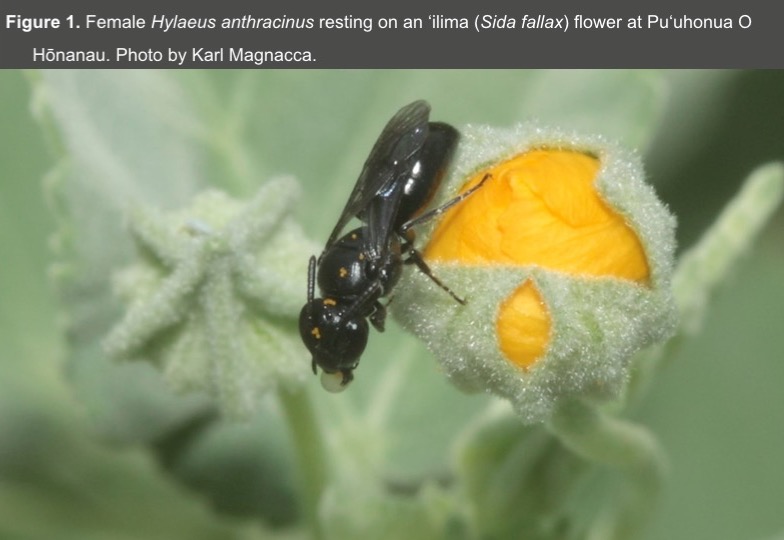
- Permits to collect Hylaeus and access state and federally managed land has been approved by the appropriate agencies (DLNR, HAVO, HALE, and PUHO) (Figure 2).
- A permit to lethally take the endangered H. anthracinus for genome sequencing and population genomic analysis is in review at the U.S. FWS (Objectives 2 and 3).
- Objective 1 is near completion and will be discussed here (Figure 2a).

- Objective 1 | Investigate non-lethal DNA sampling strategies.
- Problem: genomic DNA extraction of insects typically requires lethally sampling the individual of interest. Lethal take of an endangered species is counterproductive and may negatively impact rebounding populations. In Objective 1, we explore non-lethal tissue sampling techniques to study the conservation genetics of the endangered Hawaiian Hylaeus.
- Question 1) Is there a difference in mean survivorship of Hylaeus bees that have had either one midleg or one antennae excised from their body?
- Question 2) Is there a difference in median genomic DNA concentration (ng/uL) extracted from whole specimens, midleg, antennae, or aborted nest cells?
Objective 1 Methods
- Experimental Design
- 30 aborted nest cells were transported from O'ahu to Hilo for analysis.
- 90 Hylaeus (36 males, 54 females) were collected at two fields sites on Hawai'i Island: Pu'u O'o trail (19.68, -155.38) and Pu'u huluhulu (19.69, -155.46) in June 2018.
- Bees were transported live to UHH and kept in a 3.5 oz solo cup with 3 cotton balls and covered by a permeable membrane. One cotton ball was hydrated with 20% sugar solution ad libitum daily to feed each subject (Figure 3a).

- 30 bees were randomly selected and used as experimental controls (no tissues were excised). The remaining 60 bees were split into two groups: an antennae group and a mid-leg group (Figure 3b,c).
- Prior to dissection, bees were chilled for ~1 minute in -4 degrees C to limit motility and promote tissue removal precision.
- Antennae Group: The left antennae of 30 bees was clipped with an iris scissors as proximal to the head as possible.
- Mid-leg group: The left mid-leg of 30 bees was clipped with an iris scissors. In both procedures, the bees will be briefly chilled on ice for ~1 minute to limit mobility to promote tissue removal precision.
- Tissues were placed in -20 degrees C until needed for genomic DNA extraction.
- Control and treatment bees subjects were all placed in individual solo cup chambers (Figure 3b, c) and observed until they expired. Date of collection, dissection, and expiration were recorded.
- genomic DNA extraction
- gDNA was extracted from all excised tissues, including expired control bees (whole specimens) with a Qiagen DNeasy blood and tissue kit (Qiagen, USA).
- gDNA from 30 cells of aborted Hylaeus bee nests was extracted.
- gDNA was eluted in 50 uL of the manufacturer's elution buffer.
- gDNA was quantified with a Qubit 3 Flurometer (Thermofisher Scientific, USA) and associated reagents.
- Statistical Analyses
- We tested for the effect of the treatments (i.e., antennae, midleg) and control on mean bee survivorship with a simple one-way ANOVA
- We tested for the effect of the treatment (i.e., antennae, midleg, nestcell) and control groups on median gDNA concentration with a Kruskal-Wallis Rank Sum Test and post-hoc Dunn Test for multiple comparisons.
- Significance of the test statistic was set to an alpha of 0.05.
- Data visualization and analyses were performed on the R statistical computing platform (R Core Development Team, 2005).
Results & Future Direction
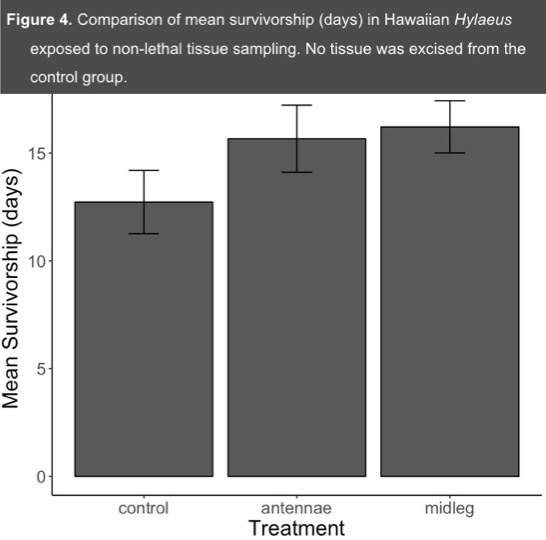
- We found no significant difference in mean survivorship across the treatment and control groups (F = 1.77, df = 2,70, P = 0.18) (Figure 4).
- We found a significant difference in median gDNA concentrations (ng/uL) across all three treatments and control groups (X2 = 83.53, df = 3, P < 0.001) (Figure 5).
- All pairwise comparisons of the four gDNA treatments are significant (P < 0.001, Mantennae = 0.15 ng/UL; Mmidleg = 0.07, Mnestcell = 7.78, Mcontrol = 25.4; M = Median) (Figure 5).
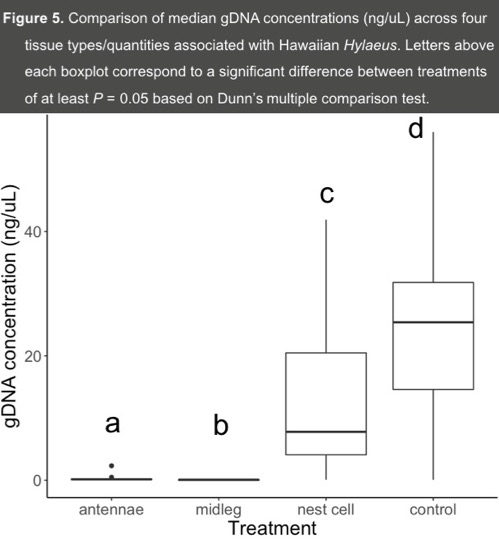
- While there is no difference in mean survivorship between excised tissue groups and the control (Figure 4), the limited tissue obtained from a non-lethal approach yields little gDNA (Figure 5).
- A preliminary assessment on the utility of non-lethally sampled gDNA for molecular analyses such as COI barcoding suggests that limited gDNA is sufficient for PCR amplificaiton.
- Future work by our group will include an assessment of non-lethally sampeled gDNA for population genomic analyses (e.g., ddRAD).
- gDNA from aborted nest cells might be a viable route to monitor populations for management and conservation purposes.
